
CHEBI:37736
| Name | D-fructofuranose 1,6-bisphosphate | 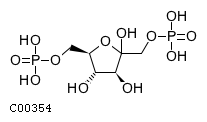 Download: mol | sdf |
| Synonyms | 01,6-di-o-phosphono-d-fructose; D-fructose 1,6-bisphosphate; D-fructose 1,6-bis(dihydrogen phosphate); | |
| Definition | The furanose form of D-fructose 1,6-bisphosphate. | |
| Molecular Weight (Exact mass) | 340.1157 (339.996) | |
| Molecular Formula | C6H14O12P2 | |
| SMILES | O[C@H]1[C@H](O)C(O)(COP(O)(O)=O)O[C@@H]1COP(O)(O)=O | |
| InChI | InChI=1S/C6H14O12P2/c7-4-3(1-16-19(10,11)12)18-6(9,5(4)8)2-17-20(13,14)15/h3-5,7-9H,1-2H2,(H2,10,11,12)(H2,13,14,15)/t3-,4-,5+,6?/m1/s1 | |
| InChI Key | RNBGYGVWRKECFJ-VRPWFDPXSA-N | |
| Crosslinking annotations | KEGG:C00354 | 3DMET:B04673 | CAS:488-69-7 | ChEBI:16905 | ChEBI:37736 | ChEMBL:CHEMBL1089962 | ChEMBL:CHEMBL2111113 | ChEMBL:CHEMBL97893 | KNApSAcK:C00007386 | NIKKAJI:J15.941G | PDB-CCD:AFP | PDB-CCD:FBP | PubChem:3647 | |
| Pathway ID | Pathway Name | Pathway Description (KEGG) |
| map00400 | Phenylalanine, tyrosine and tryptophan biosynthesis | NA |
| map00680 | Methane metabolism | Methane is metabolized principally by methanotrophs and methanogens in the global carbon cycle. Methanotrophs consume methane as the only source of carbon, while methanogens produce methane as a metabolic byproduct. Methylotrophs, which are microorganisms that can obtain energy for growth by oxidizing one-carbon compounds, such as methanol and methane, are situated between methanotrophs and methanogens. Methanogens can obtain energy for growth by converting a limited number of substrates to methane under anaerobic conditions. Three types of methanogenic pathways are known: CO2 to methane [MD:M00567], methanol to methane [MD:M00356], and acetate to methane [MD:M00357]. Methanogens use 2-mercaptoethanesulfonate (CoM; coenzyme M) as the terminal methyl carrier in methanogenesis and have four enzymes for CoM biosynthesis [MD:M00358]. Coenzyme B-Coenzyme M heterodisulfide reductase (Hdr), requiring for the final reaction steps of methanogenic pathway, is divided into two types: cytoplasmic HdrABC in most methanogens and membrane-bound HdrED in Methanosarcina species. In methanotrophs and methyltrophs methane is oxidized to form formaldehyde, which is at the diverging point for further oxidation to CO2 for energy source and assimilation for biosynthesis. There are three pathways that convert formaldehyde to C2 or C3 compounds: serine pathway [MD:M00346], ribulose monophosphate pathway [MD:M00345], and xylulose monophosphate pathway [MD:M00344]. The first two pathways are found in prokaryotes and the third is found in yeast. As a special case of methylotrophs, various amines can be used as carbon sources in trimethylamine metabolism [MD:M00563]. |
| map00710 | Carbon fixation in photosynthetic organisms | NA |
| map01100 | Metabolic pathways | NA |
| map01110 | Biosynthesis of secondary metabolites | NA |
| map01120 | Microbial metabolism in diverse environments | NA |
| map01130 | Biosynthesis of antibiotics | NA |
| map01200 | Carbon metabolism | Carbon metabolism is the most basic aspect of life. This map presents an overall view of central carbon metabolism, where the number of carbons is shown for each compound denoted by a circle, excluding a cofactor (CoA, CoM, THF, or THMPT) that is replaced by an asterisk. The map contains carbon utilization pathways of glycolysis (map00010), pentose phosphate pathway (map00030), and citrate cycle (map00020), and six known carbon fixation pathways (map00710 and map00720) as well as some pathways of methane metabolism (map00680). The six carbon fixation pathways are: (1) reductive pentose phosphate cycle (Calvin cycle) in plants and cyanobacteria that perform oxygenic photosynthesis, (2) reductive citrate cycle in photosynthetic green sulfur bacteria and some chemolithoautotrophs, (3) 3-hydroxypropionate bi-cycle in photosynthetic green nonsulfur bacteria, two variants of 4-hydroxybutyrate pathways in Crenarchaeota called (4) hydroxypropionate-hydroxybutyrate cycle and (5) dicarboxylate-hydroxybutyrate cycle, and (6) reductive acetyl-CoA pathway in methanogenic bacteria. |
| map04152 | AMPK signaling pathway | AMP-activated protein kinase (AMPK) is a serine threonine kinase that is highly conserved through evolution. AMPK system acts as a sensor of cellular energy status. It is activated by increases in the cellular AMP:ATP ratio caused by metabolic stresses that either interfere with ATP production (eg, deprivation for glucose or oxygen) or that accelerate ATP consumption (eg, muscle contraction). Several upstream kinases, including liver kinase B1 (LKB1), calcium/calmodulin kinase kinase-beta (CaMKK beta), and TGF-beta-activated kinase-1 (TAK-1), can activate AMPK by phosphorylating a threonine residue on its catalytic alpha-subunit. Once activated, AMPK leads to a concomitant inhibition of energy-consuming biosynthetic pathways, such as protein, fatty acid and glycogen synthesis, and activation of ATP-producing catabolic pathways, such as fatty acid oxidation and glycolysis. |
| map04922 | Glucagon signaling pathway | Glucagon is conventionally regarded as a counterregulatory hormone for insulin and plays a critical anti-hypoglycemic role by maintaining glucose homeostasis in both animals and humans. To increase blood glucose, glucagon promotes hepatic glucose output by increasing glycogenolysis and gluconeogenesis and by decreasing glycogenesis and glycolysis in a concerted fashion via multiple mechanisms. Glucagon also stimulates hepatic mitochondrial beta-oxidation to supply energy for glucose production. Glucagon performs its main effect via activation of adenylate cyclase. The adenylate-cyclase-derived cAMP activates protein kinase A (PKA), which then phosphorylates downstream targets, such as cAMP response element binding protein (CREB) and the bifunctional enzyme 6-phosphofructo-2-kinase/ fructose-2,6-bisphosphatase (one of the isoforms being PFK/FBPase 1, encoded by PFKFB1). |
| map05230 | Central carbon metabolism in cancer | Malignant transformation of cells requires specific adaptations of cellular metabolism to support growth and survival. In the early twentieth century, Otto Warburg established that there are fundamental differences in the central metabolic pathways operating in malignant tissue. He showed that cancer cells consume a large amount of glucose, maintain high rate of glycolysis and convert a majority of glucose into lactic acid even under normal oxygen concentrations (Warburg's Effects). More recently, it has been recognized that the 'Warburg effect' encompasses a similarly increased utilization of glutamine. From the intermediate molecules provided by enhanced glycolysis and glutaminolysis, cancer cells synthesize most of the macromolecules required for the duplication of their biomass and genome. These cancer-specific alterations represent a major consequence of genetic mutations and the ensuing changes of signalling pathways in cancer cells. Three transcription factors, c-MYC, HIF-1 and p53, are key regulators and coordinate regulation of cancer metabolism in different ways, and many other oncogenes and tumor suppressor genes cluster along the signaling pathways that regulate c-MYC, HIF-1 and p53. |

