Top
PLPPR3
| Gene Name | PLPPR3 (QuickGO) | (A quick tutorial to explore the interctive visulaization) |
| Synonyms | PLPPR3 , LPPR3 | |
| Protein Name | PLPPR3 |
|
| Alternative Name(s) |
Phospholipid phosphatase-related protein type 3;3.1.3.4;Lipid phosphate phosphatase-related protein type 3;PAP-2-like protein 2;Plasticity-related gene 2 protein;PRG-2; | |
| EntrezGene ID | 79948 (Comparitive Toxicogenomics) | |
| UniProt AC (Human) | Q6T4P5 (protein sequence) | |
| Enzyme Class | EC 3.1.3.4 (BRENDA | |
| Molecular Weight | 76037 Dalton | |
| Protein Length | 718 amino acids (AA) | |
| Genome Browsers | NCBI | ENSG00000129951 (Ensembl) | UCSC | |
| Crosslinking annotations | Query our ID-mapping table | |
| Orthologues | Quest For Orthologues (QFO) | GeneTree | |
| Classification | Superfamily: glucose_lipid phosphatases | Historic class: Phosphatidic acid phosphatase (PAP) | CATH ID: 1.20.144.10 | SCOP Fold: Chloroperoxidase | |
| Phosphatase activity | active | Catalytic signature motif: unknown | |
| Domain organization, Expression, Diseases | (show / hide) |
| Protein Domain | InterPro | Pfam | SMART | 3D Structure | PDB | DrugPort | ModBase | SwissModel |
| Polypeptide organization (Pfam) | No Pfam domain found | ||
| Catalytic Site | Mechanism and Catalytic Site Atlas | ||
| Gene Expression | Gene Expression Atlas | Diseases | OMIM | ClinVar |
| Protein-protein interactions | STRING | IntAct | MINT | BioGRID | ||
| Phosphosites | Phospho.ELM | PhosphoSitePlus | ||
| Localization, Function, Catalytic activity and Sequence | (show / hide) |
| Localization (UniProt annotation) | |||
| Membrane | |||
| Function (UniProt annotation) | |||
| NA | |||
| Catalytic Activity (UniProt annotation) | |||
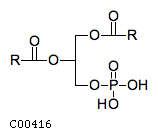
| A 1,2-diacylglycerol 3-phosphate + H(2)O = a1,2-diacyl-sn-glycerol + phosphate | |||
| Protein Sequence | |||
| MISTKEKNKIPKDSMTLLPCFYFVELPIVASSIVSLYFLELTDLFKPAKVGFQCYDRTLSMPYVETNEELIPLLMLLSLA FAAPAASIMVAEGMLYCLQSRLWGRAGGPAGAEGSINAGGCNFNSFLRRTVRFVGVHVFGLCATALVTDVIQLATGYHTP FFLTVCKPNYTLLGTSCEVNPYITQDICSGHDIHAILSARKTFPSQHATLSAFAAVYVSMYFNSVISDTTKLLKPILVFA FAIAAGVCGLTQITQYRSHPVDVYAGFLIGAGIAAYLACHAVGNFQAPPAEKPAAPAPAKDALRALTQRGHDSVYQQNKS VSTDELGPPGRLEGAPRPVAREKTSLGSLKRASVDVDLLAPRSPMAKENMVTFSHTLPRASAPSLDDPARRHMTIHVPLD ASRSKQLISEWKQKSLEGRGLGLPDDASPGHLRAPAEPMAEEEEEEEDEEEEEEEEEEEDEGPAPPSLYPTVQARPGLGP RVILPPRAGPPPLVHIPEEGAQTGAGLSPKSGAGVRAKWLMMAEKSGAAVANPPRLLQVIAMSKAPGAPGPKAAETASSS SASSDSSQYRSPSDRDSASIVTIDAHAPHHPVVHLSAGGAPWEWKAAGGGAKAEADGGYELGDLARGFRGGAKPPGVSPG SSVSDVDQEEPRFGAVATVNLATGEGLPPLGAADGALGPGSRESTLRRHAGGLGLAEREAEAEAEGYFRKMQARRFPD |
| Motif information from Eukaryotic Linear Motif atlas (ELM) | (show / hide) |
| Gene Ontology (P: Process; F: Function and C: Component terms) | (show / hide) | |
| GO:0005887 | C:integral component of plasma membrane |
| GO:0006644 | P:phospholipid metabolic process |
| GO:0007165 | P:signal transduction |
| GO:0008195 | F:phosphatidate phosphatase activity |
| GO:0042577 | F:lipid phosphatase activity |
| GO:0046839 | P:phospholipid dephosphorylation |