Top
PTPMT1
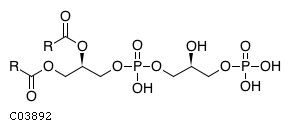
Pathway ID Pathway Name Pathway Description (KEGG) R-HSA-1483148 Synthesis of PG. Phosphatidylglycerol (PG) is synthesised at the inner mitochondrial (IM) membrane, phosphatidic acid (PA) and cytidine triphosphate (CTP) are converted into cytidine diphosphate-diacylglycerol (CDP-DAG), which in turn is converted with glycerol-3-phosphate (G3P) into phosphatidylglycerophosphate (PGP) and cytidine monophosphate (CMP). Finally, PGP is dephosphorylated to PG. In addition, PG can be synthesised at the endoplamic reticulum (ER) membrane when phospholipase D transphosphatidylates phosphatidylcholine (PC) with glycerol to displace choline (Cho) and form PG (Piazza & Marmer 2007, Stuhne-Sekalec et al. 1986, Lykidis et al. 1997, Cao & Hatch 1994)
Visualize Download
Interacting partner Detection method Interaction type PubMed IDs Interaction Score Source AFG3L2 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID ATAD3A Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID ATP2A2 Affinity Capture-MS physical 27880917 , (Europe PMC )NA BioGRID ATP5F1C Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID ATP5F1E Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID ATP5MD Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID ATP5PB Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID ATP5PD Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID ATP5PF Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID ATP5PO Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID BPNT1 Affinity Capture-MS physical 28514442 , (Europe PMC )NA BioGRID BSG Affinity Capture-MS physical 26186194 , (Europe PMC )NA BioGRID C1QBP Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID CBWD1 Affinity Capture-MS physical 26186194 , 28514442 , (Europe PMC )NA BioGRID CD79A Affinity Capture-MS physical 28514442 , (Europe PMC )NA BioGRID CHCHD2 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID CISD1 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID CLPB Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID CLPX Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID COPS5 Affinity Capture-MS physical 21145461 , (Europe PMC )NA BioGRID COQ5 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID COQ7 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID COQ9 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID COX5A Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID COX7A2 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID CPT2 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID CYCS Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID DUSP6 Affinity Capture-MS physical 27432908 , (Europe PMC )NA BioGRID ECH1 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID ERGIC3 Affinity Capture-MS physical 26186194 , (Europe PMC )NA BioGRID FBXO6 Affinity Capture-MS physical 22268729 , (Europe PMC )NA BioGRID FKBP8 Affinity Capture-MS physical 27880917 , (Europe PMC )NA BioGRID GLS Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID HINT3 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID IL13RA2 Affinity Capture-MS physical 28514442 , (Europe PMC )NA BioGRID IMMT Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID IPO9 Affinity Capture-MS physical 27880917 , (Europe PMC )NA BioGRID KRT31 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 25416956 , Europe PMC )0.70 BioGRID, IntAct KRT40 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 25416956 , (Europe PMC )0.56 BioGRID, IntAct KRTAP10-3 Two-hybrid physical 25416956 , (Europe PMC )NA BioGRID KRTAP10-7 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 25416956 , Europe PMC )0.49, 0.56 BioGRID, IntAct KRTAP10-8 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 25416956 , Europe PMC )0.70 BioGRID, IntAct KRTAP2-4 Two-hybrid physical 25416956 , (Europe PMC )NA BioGRID KRTAP5-9 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 25416956 , Europe PMC )0.70 BioGRID, IntAct KRTAP9-2 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 25416956 , Europe PMC )0.70 BioGRID, IntAct LAMP2 Affinity Capture-MS physical 28514442 , (Europe PMC )NA BioGRID LETM1 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID MAPK6 Two-hybrid, two hybrid physical, physical association 21900206 , (Europe PMC )0.37 BioGRID, IntAct, MINT MDFI Two-hybrid, two hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 21516116 , 25416956 , Europe PMC )0.76 BioGRID, IntAct, MINT MRPL12 Affinity Capture-MS physical 28514442 , (Europe PMC )NA BioGRID MTCH2 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID MTUS2 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 25416956 , (Europe PMC )0.56 BioGRID, IntAct NDUFA8 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID NDUFAF2 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID NDUFS3 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID NIPSNAP1 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID NOTCH2NLA Two-hybrid physical 25416956 , (Europe PMC )NA BioGRID NUP133 Affinity Capture-MS physical 27880917 , (Europe PMC )NA BioGRID OXA1L Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID OXCT1 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID PAK5 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID PAM16 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID PMPCB Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID POLDIP2 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID PTPMT1 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID RAB5IF Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID RGS20 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 21516116 , 25416956 , Europe PMC )0.70 BioGRID, IntAct SFXN1 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID SIAH1 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 25416956 , (Europe PMC )0.56 BioGRID, IntAct SLC16A1 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID SLC18A1 Affinity Capture-MS physical 28514442 , (Europe PMC )NA BioGRID SLC25A1 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID SLC25A3 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID SLC2A12 Affinity Capture-MS physical 28514442 , (Europe PMC )NA BioGRID SNAP29 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID STOML2 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID TAB1 Two-hybrid, two hybrid physical, physical association 21900206 , (Europe PMC )0.37 BioGRID, IntAct, MINT TIMM44 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID TIMM50 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID TRIM27 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 25416956 , (Europe PMC )0.56 BioGRID, IntAct UQCRB Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID UQCRC2 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID UQCRFS1 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID XPO4 Affinity Capture-MS physical 27880917 , (Europe PMC )NA BioGRID
Visualize Download
Interacting partner Detection method Interaction type PubMed IDs Interaction Score Source KRT31 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 25416956 , Europe PMC )0.70 BioGRID, IntAct KRT40 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 25416956 , (Europe PMC )0.56 BioGRID, IntAct KRTAP10-7 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 25416956 , Europe PMC )0.49, 0.56 BioGRID, IntAct KRTAP10-8 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 25416956 , Europe PMC )0.70 BioGRID, IntAct KRTAP2-3 two hybrid array, two hybrid prey pooling approach, validated two hybrid physical association 25416956 , (Europe PMC )0.56 IntAct KRTAP5-9 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 25416956 , Europe PMC )0.70 BioGRID, IntAct KRTAP9-2 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 25416956 , Europe PMC )0.70 BioGRID, IntAct MAPK6 Two-hybrid, two hybrid physical, physical association 21900206 , (Europe PMC )0.37 BioGRID, IntAct, MINT MDFI Two-hybrid, two hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 21516116 , 25416956 , Europe PMC )0.76 BioGRID, IntAct, MINT MTUS2 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 25416956 , (Europe PMC )0.56 BioGRID, IntAct NOTCH2NL two hybrid array, two hybrid prey pooling approach, validated two hybrid physical association 25416956 , Europe PMC )0.70 IntAct RGS20 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 21516116 , 25416956 , Europe PMC )0.70 BioGRID, IntAct SIAH1 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 25416956 , (Europe PMC )0.56 BioGRID, IntAct TAB1 Two-hybrid, two hybrid physical, physical association 21900206 , (Europe PMC )0.37 BioGRID, IntAct, MINT TRIM27 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 25416956 , (Europe PMC )0.56 BioGRID, IntAct
Visualize Download
Interacting partner Detection method Interaction type PubMed IDs Interaction Score Source MDFI Two-hybrid, two hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 21516116 , 25416956 , Europe PMC )0.76 BioGRID, IntAct, MINT TAB1 Two-hybrid, two hybrid physical, physical association 21900206 , (Europe PMC )0.37 BioGRID, IntAct, MINT
Visualize Download
Interacting partner Detection method Interaction type PubMed IDs Interaction Score Source AFG3L2 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID ATAD3A Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID ATP2A2 Affinity Capture-MS physical 27880917 , (Europe PMC )NA BioGRID ATP5F1C Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID ATP5F1E Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID ATP5MD Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID ATP5PB Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID ATP5PD Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID ATP5PF Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID ATP5PO Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID BPNT1 Affinity Capture-MS physical 28514442 , (Europe PMC )NA BioGRID BSG Affinity Capture-MS physical 26186194 , (Europe PMC )NA BioGRID C1QBP Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID CBWD1 Affinity Capture-MS physical 26186194 , 28514442 , (Europe PMC )NA BioGRID CD79A Affinity Capture-MS physical 28514442 , (Europe PMC )NA BioGRID CHCHD2 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID CISD1 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID CLPB Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID CLPX Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID COPS5 Affinity Capture-MS physical 21145461 , (Europe PMC )NA BioGRID COQ5 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID COQ7 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID COQ9 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID COX5A Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID COX7A2 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID CPT2 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID CYCS Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID DUSP6 Affinity Capture-MS physical 27432908 , (Europe PMC )NA BioGRID EBI-10171679 two hybrid array, two hybrid prey pooling approach, validated two hybrid physical association 25416956 , (Europe PMC )0.56 IntAct ECH1 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID ERGIC3 Affinity Capture-MS physical 26186194 , (Europe PMC )NA BioGRID FBXO6 Affinity Capture-MS physical 22268729 , (Europe PMC )NA BioGRID FKBP8 Affinity Capture-MS physical 27880917 , (Europe PMC )NA BioGRID GLS Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID HINT3 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID IL13RA2 Affinity Capture-MS physical 28514442 , (Europe PMC )NA BioGRID IMMT Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID IPO9 Affinity Capture-MS physical 27880917 , (Europe PMC )NA BioGRID KRT31 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 25416956 , Europe PMC )0.70 BioGRID, IntAct KRT40 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 25416956 , (Europe PMC )0.56 BioGRID, IntAct KRTAP10-3 Two-hybrid physical 25416956 , (Europe PMC )NA BioGRID KRTAP10-7 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 25416956 , Europe PMC )0.49, 0.56 BioGRID, IntAct KRTAP10-8 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 25416956 , Europe PMC )0.70 BioGRID, IntAct KRTAP2-3 two hybrid array, two hybrid prey pooling approach, validated two hybrid physical association 25416956 , (Europe PMC )0.56 IntAct KRTAP2-4 Two-hybrid physical 25416956 , (Europe PMC )NA BioGRID KRTAP5-9 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 25416956 , Europe PMC )0.70 BioGRID, IntAct KRTAP9-2 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 25416956 , Europe PMC )0.70 BioGRID, IntAct LAMP2 Affinity Capture-MS physical 28514442 , (Europe PMC )NA BioGRID LETM1 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID MAPK6 Two-hybrid, two hybrid physical, physical association 21900206 , (Europe PMC )0.37 BioGRID, IntAct, MINT MDFI Two-hybrid, two hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 21516116 , 25416956 , Europe PMC )0.76 BioGRID, IntAct, MINT MRPL12 Affinity Capture-MS physical 28514442 , (Europe PMC )NA BioGRID MTCH2 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID MTUS2 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 25416956 , (Europe PMC )0.56 BioGRID, IntAct NDUFA8 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID NDUFAF2 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID NDUFS3 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID NIPSNAP1 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID NOTCH2NL two hybrid array, two hybrid prey pooling approach, validated two hybrid physical association 25416956 , Europe PMC )0.70 IntAct NOTCH2NLA Two-hybrid physical 25416956 , (Europe PMC )NA BioGRID NUP133 Affinity Capture-MS physical 27880917 , (Europe PMC )NA BioGRID OXA1L Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID OXCT1 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID PAK5 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID PAM16 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID PMPCB Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID POLDIP2 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID PTPMT1 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID RAB5IF Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID RGS20 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 21516116 , 25416956 , Europe PMC )0.70 BioGRID, IntAct SFXN1 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID SIAH1 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 25416956 , (Europe PMC )0.56 BioGRID, IntAct SLC16A1 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID SLC18A1 Affinity Capture-MS physical 28514442 , (Europe PMC )NA BioGRID SLC25A1 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID SLC25A3 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID SLC2A12 Affinity Capture-MS physical 28514442 , (Europe PMC )NA BioGRID SNAP29 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID STOML2 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID TAB1 Two-hybrid, two hybrid physical, physical association 21900206 , (Europe PMC )0.37 BioGRID, IntAct, MINT TIMM44 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID TIMM50 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID TRIM27 Two-hybrid, two hybrid array, two hybrid prey pooling approach, validated two hybrid physical, physical association 25416956 , (Europe PMC )0.56 BioGRID, IntAct UQCRB Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID UQCRC2 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID UQCRFS1 Affinity Capture-MS physical 27499296 , (Europe PMC )NA BioGRID XPO4 Affinity Capture-MS physical 27880917 , (Europe PMC )NA BioGRID