
CHEBI:15811
| Name | O-phospho-L-serine | 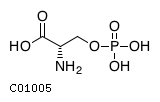 Download: mol | sdf |
| Synonyms | 0(+)-l-serine dihydrogen phosphate (ester); (2s)-2-amino-3-(phosphonooxy)propanoic acid; (2s)-2-azaniumyl-3-(phosphonatooxy)propanoate; (s)-2-amino-3-hydroxypropanoic acid 3-phosphate; 3-phosphoserine; Dexfosfoserine; L-o-phosphoserine; O-phospho-l-serine; O-phosphoserine; Phosphoserine; | |
| Definition | The L-enantiomer of O-phosphoserine. | |
| Molecular Weight (Exact mass) | 185.0725 (185.0089) | |
| Molecular Formula | C3H8NO6P | |
| SMILES | N[C@@H](COP(O)(O)=O)C(O)=O | |
| InChI | InChI=1S/C3H8NO6P/c4-2(3(5)6)1-10-11(7,8)9/h2H,1,4H2,(H,5,6)(H2,7,8,9)/t2-/m0/s1 | |
| InChI Key | BZQFBWGGLXLEPQ-REOHCLBHSA-N | |
| Crosslinking annotations | KEGG:C01005 | 3DMET:B01366 | CAS:407-41-0 | ChEBI:15811 | ChEMBL:CHEMBL284377 | KNApSAcK:C00007287 | NIKKAJI:J136.545B | PDB-CCD:SEP | PubChem:4251 | |
| Pathway ID | Pathway Name | Pathway Description (KEGG) |
| map00260 | Glycine, serine and threonine metabolism | Serine is derived from 3-phospho-D-glycerate, an intermediate of glycolysis [MD:M00020], and glycine is derived from serine. Threonine is an essential amino acid, which animals cannot synthesize. In bacteria and plants, threonine is derived from aspartate [MD:M00018]. |
| map00270 | Cysteine and methionine metabolism | Cysteine and methionine are sulfur-containing amino acids. Cysteine is synthesized from serine through different pathways in different organism groups. In bacteria and plants, cysteine is converted from serine (via acetylserine) by transfer of hydrogen sulfide [MD:M00021]. In animals, methionine-derived homocysteine is used as sulfur source and its condensation product with serine (cystathionine) is converted to cysteine [MD:M00338]. Cysteine is metabolized to pyruvate in multiple routes. Methionine is an essential amino acid, which animals cannot synthesize. In bacteria and plants, methionine is synthesized from aspartate [MD:M00017]. S-Adenosylmethionine (SAM), synthesized from methionine and ATP, is a methyl group donor in many important transfer reactions including DNA methylation for regulation of gene expression. SAM may also be used to regenerate methionine in the methionine salvage pathway [MD:M00034]. |
| map00680 | Methane metabolism | Methane is metabolized principally by methanotrophs and methanogens in the global carbon cycle. Methanotrophs consume methane as the only source of carbon, while methanogens produce methane as a metabolic byproduct. Methylotrophs, which are microorganisms that can obtain energy for growth by oxidizing one-carbon compounds, such as methanol and methane, are situated between methanotrophs and methanogens. Methanogens can obtain energy for growth by converting a limited number of substrates to methane under anaerobic conditions. Three types of methanogenic pathways are known: CO2 to methane [MD:M00567], methanol to methane [MD:M00356], and acetate to methane [MD:M00357]. Methanogens use 2-mercaptoethanesulfonate (CoM; coenzyme M) as the terminal methyl carrier in methanogenesis and have four enzymes for CoM biosynthesis [MD:M00358]. Coenzyme B-Coenzyme M heterodisulfide reductase (Hdr), requiring for the final reaction steps of methanogenic pathway, is divided into two types: cytoplasmic HdrABC in most methanogens and membrane-bound HdrED in Methanosarcina species. In methanotrophs and methyltrophs methane is oxidized to form formaldehyde, which is at the diverging point for further oxidation to CO2 for energy source and assimilation for biosynthesis. There are three pathways that convert formaldehyde to C2 or C3 compounds: serine pathway [MD:M00346], ribulose monophosphate pathway [MD:M00345], and xylulose monophosphate pathway [MD:M00344]. The first two pathways are found in prokaryotes and the third is found in yeast. As a special case of methylotrophs, various amines can be used as carbon sources in trimethylamine metabolism [MD:M00563]. |
| map00970 | Aminoacyl-tRNA biosynthesis | NA |
| map01100 | Metabolic pathways | NA |
| map01120 | Microbial metabolism in diverse environments | NA |
| map01130 | Biosynthesis of antibiotics | NA |
| map01200 | Carbon metabolism | Carbon metabolism is the most basic aspect of life. This map presents an overall view of central carbon metabolism, where the number of carbons is shown for each compound denoted by a circle, excluding a cofactor (CoA, CoM, THF, or THMPT) that is replaced by an asterisk. The map contains carbon utilization pathways of glycolysis (map00010), pentose phosphate pathway (map00030), and citrate cycle (map00020), and six known carbon fixation pathways (map00710 and map00720) as well as some pathways of methane metabolism (map00680). The six carbon fixation pathways are: (1) reductive pentose phosphate cycle (Calvin cycle) in plants and cyanobacteria that perform oxygenic photosynthesis, (2) reductive citrate cycle in photosynthetic green sulfur bacteria and some chemolithoautotrophs, (3) 3-hydroxypropionate bi-cycle in photosynthetic green nonsulfur bacteria, two variants of 4-hydroxybutyrate pathways in Crenarchaeota called (4) hydroxypropionate-hydroxybutyrate cycle and (5) dicarboxylate-hydroxybutyrate cycle, and (6) reductive acetyl-CoA pathway in methanogenic bacteria. |
| map01230 | Biosynthesis of amino acids | This map presents a modular architecture of the biosynthesis pathways of twenty amino acids, which may be viewed as consisting of the core part and its extensions. The core part is the KEGG module for conversion of three-carbon compounds from glyceraldehyde-3P to pyruvate [MD:M00002], together with the pathways around serine and glycine. This KEGG module is the most conserved one in the KEGG MODULE database and is found in almost all the completely sequenced genomes. The extensions are the pathways containing the reaction modules RM001, RM033, RM032, and RM002 for biosynthesis of branched-chain amino acids (left) and basic amino acids (bottom), and the pathways for biosynthesis of histidine and aromatic amino acids (top right). It is interesting to note that the so-called essential amino acids that cannot be synthesized in human and other organisms generally appear in these extensions. Furthermore, the bottom extension of basic amino acids appears to be most divergent containing multiple pathways for lysine biosynthesis and multiple gene sets for arginine biosynthesis. |

